UF Health researchers’ new findings have implications for muscle development, disease
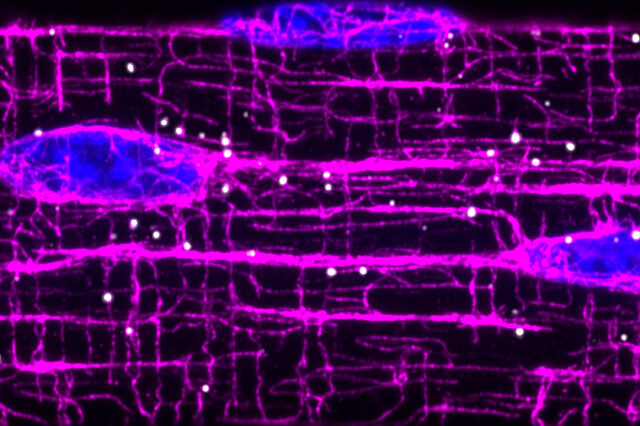
To allow movement and function, our muscles use tiny “contraction machines” inside of cells. Each on its own is too weak to move even a fingernail, but muscles can move entire bodies through precise organization and coordination of these machines.
Now a group of University of Florida Health researchers has learned more about how skeletal muscle cells organize their components to function properly and sustain life. The findings have important implications for muscle development, maintenance and disease, the researchers said.
As the largest cell in the human body, muscle cells need to transport crucial cellular products across a comparatively long distance. Understanding how that process — essentially an “intracellular highway system” — works is an key step in better understanding muscle diseases, said Lance T. Denes, Ph.D., a co-author of the findings published Oct. 27 in the journal Nature Communications.
The research team focused on how cargos inside of muscle cells are transported from a “command center” — the nucleus — to more distant outposts. One of those molecules, RNA, plays a key role in converting genetic information into proteins the body needs.
In muscle cells, the machines that allow movement are built from hundreds of proteins, each fitting together like an intricate puzzle. RNA molecules contain the instructions for building and maintaining these machines and need to be delivered to the appropriate “construction sites” for muscle cells to function. In their work, the researchers show that an elaborate system of filaments called microtubules function as “highways” and direct the transport of RNA molecules throughout cell, thus allowing production of proteins “on site.”
The results are important because most of the genes involved in this transport pathway are implicated in a variety of muscle diseases, including Duchenne muscular dystrophy (a progressive muscle-wasting disorder) and myotonic dystrophy (a genetic disorder that causes muscle weakness and atrophy).
To establish their findings, the researchers developed a method to detect single RNA molecules and protein markers in skeletal muscles as well as computational methods to characterize their positions. By disrupting the function of microtubules, they found that RNAs began to aggregate around the nucleus, causing an intracellular traffic jam. This work opens up an intriguing link between the effectiveness of RNA transport and the aggregation of disease proteins.
“This study will inspire future work to assess the impact of altered RNA transport on disease origin and development. This work suggests that paying closer attention to the locations and mechanisms of travel for RNA is critical for understanding complex biological systems such as large, differentiated cells,” said Denes, who is now a postdoctoral associate at New York University.
Next, the researchers want to know how much RNA transport is affected in many muscle diseases. That would involve using the technology they developed to visualize RNA molecules in mice and diseased human tissue, Denes said.
The research team was led by Eric T. Wang, an associate professor in the UF College of Medicine’s department of molecular genetics and microbiology. It also included Chase P. Kelley, a doctoral candidate in Wang’s lab. Research funding was provided by the National Institutes of Health and the Chan-Zuckerberg Initiative.
About the author
